Arthroderma
The genus Arthroderma currently encompasses 27 species (Hainsworth et al. 2021), mostly occurring in soil, cave environments and infrequently from human and animal cutaneous infections. They are all geophilic dermatophyte species which play an important ecological role in the decomposition of keratin from animal hair and skin.
The molecular taxonomy of this genus has also been ambiguous with de Hoog et al. (2017), transferring Trichophyton terrestre and T. ajelloi to Arthroderma as A. insingulare and A. uncinatum respectively. However, a more recent multi-gene phylogenetic study by Hainsworth et al. (2021) has validated Arthroderma terrestre as the correct name for Trichophyton terrestre.
Morphological description: Colonies mostly granular to cottony, yellowish to brownish, with a cream-coloured or brown colony reverse. Macroconidia are multicelled, (sub)hyaline, clavate, fusiform or cigar-shaped and are thick- and rough-walled. Microconidia are hyaline, 1-celled, clavate and smooth-walled.
Species descriptions
-
Arthroderma terrestre
Synonymy:
Trichophyton terrestreArthroderma insingulare is a geophilic fungus of worldwide distribution which may occur as a saprophytic contaminant on humans and animals. Durie and Frey (1957) first described this soil fungus as Trichophyton terrestre from New South Wales, Australia. Since then T. terrestre has been described as an anamorph of three different species of Arthroderma; A. insingulare, A. lenticulare and A. quadrifidum (Padhye and Carmichael, 1972). However, ITS and D1/D2 sequencing of the original isolates has now identified this fungus as Arthroderma terrestre (Hainsworth et al. 2021).
Arthroderma terrestre is the most frequently encountered Arthroderma species recovered from human and animal skin and hair specimens.
RG-1 organism.
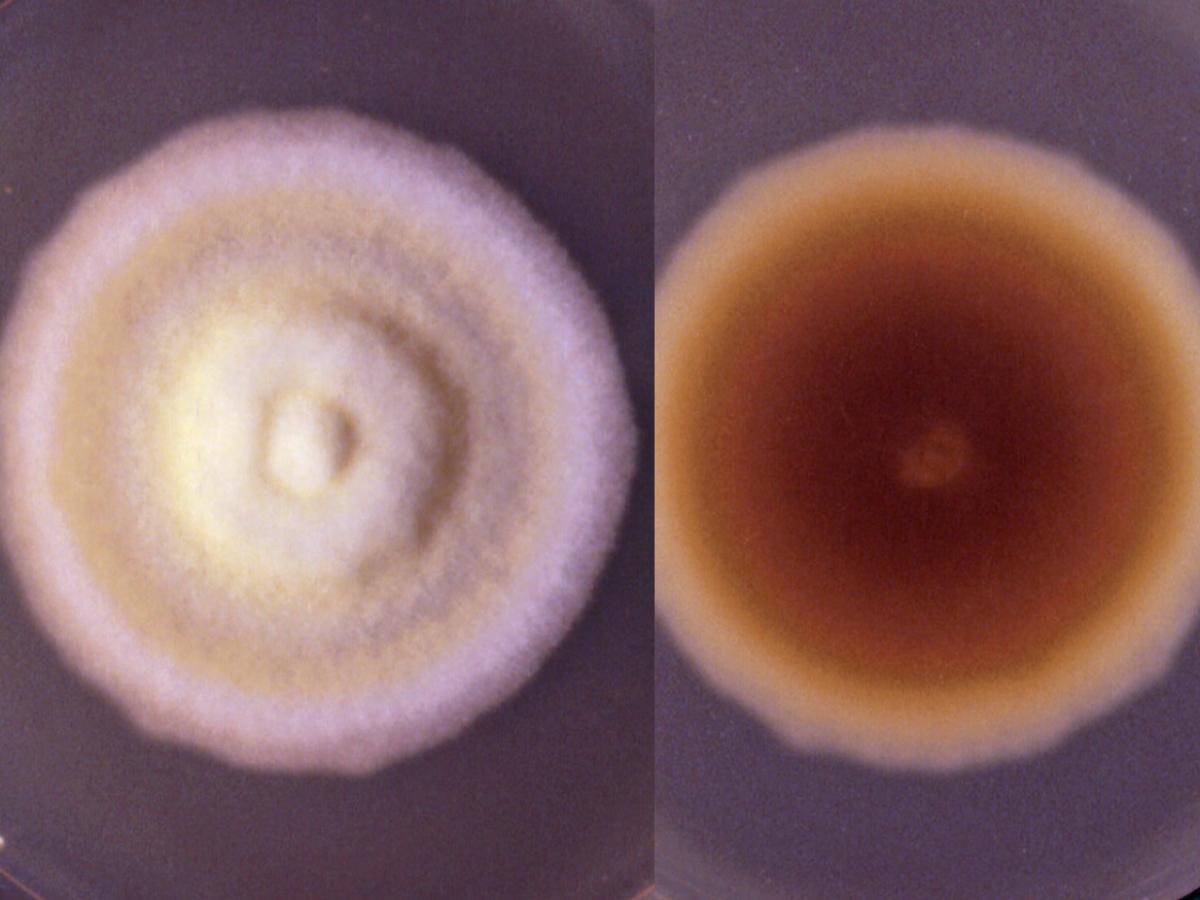
Arthroderma insingulare culture.
Morphological description:
Colonies are usually flat to downy with a suede-like to granular texture resembling T. mentagrophytes. The surface colour may range from white to cream, buff to yellow, or greenish-yellow. Reverse pigmentation is usually yellowish-brown although some variants have a deep rose red reverse. Microconidia are large, clavate or pedicellate, usually exhibiting transition forms to more or less abundant lateral macroconidia. Macroconidia are clavate to cylindrical with rounded ends, smooth and thin-walled, and are two to six-celled. Chlamydospores, hyphal spirals, racquet mycelium and antler hyphae may also be present. No growth at 37C.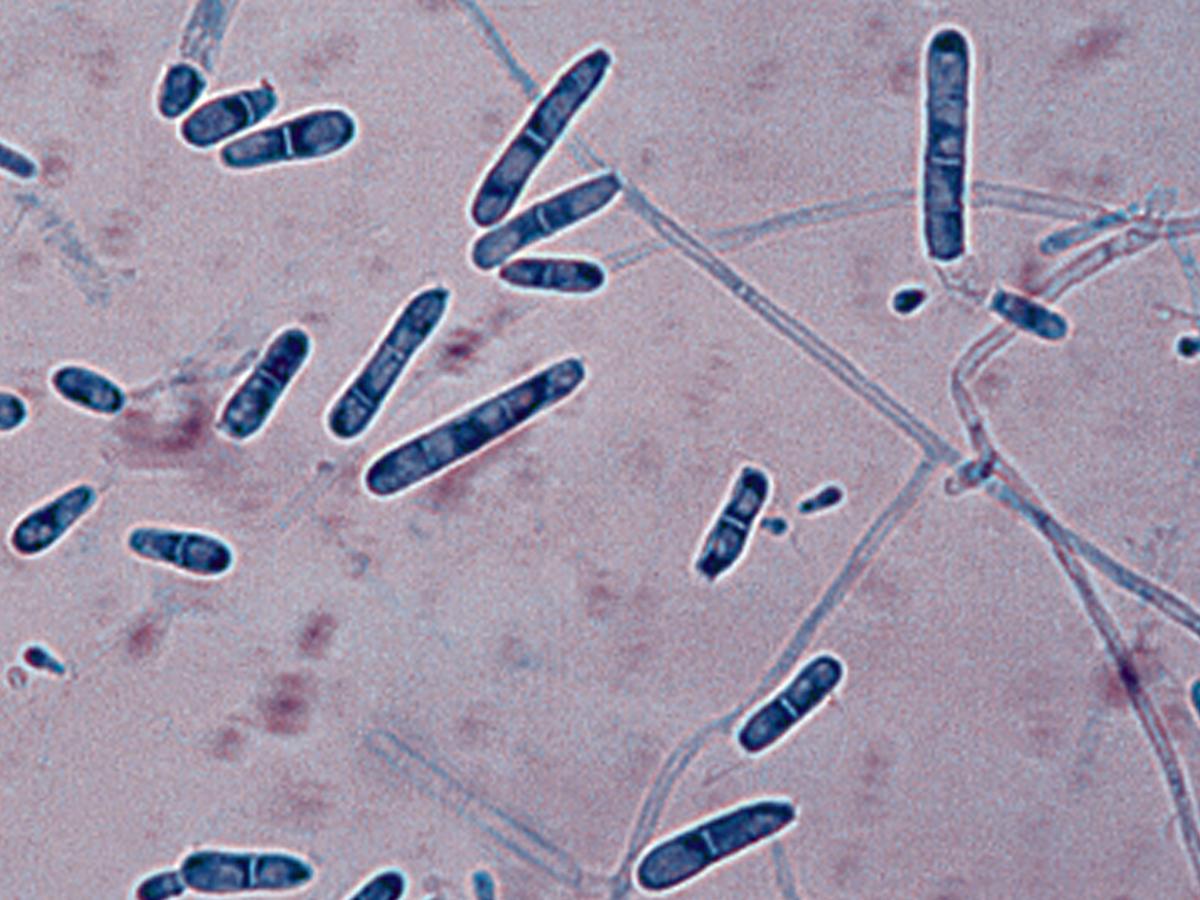
Arthroderma insingulare macroconidia.
Molecular identification:
ITS and D1/D2 sequencing is recommended for definitive identification of isolates. -
Arthroderma uncinatum
Synonymy:
Trichophyton ajelloiArthroderma uncinatum is a geophilic fungus with a worldwide distribution which may occur as a saprophytic contaminant on humans and animals but infections are doubtful. Not known to invade hair in vivo, but produces hair perforations in vitro.
RG-1 organism.
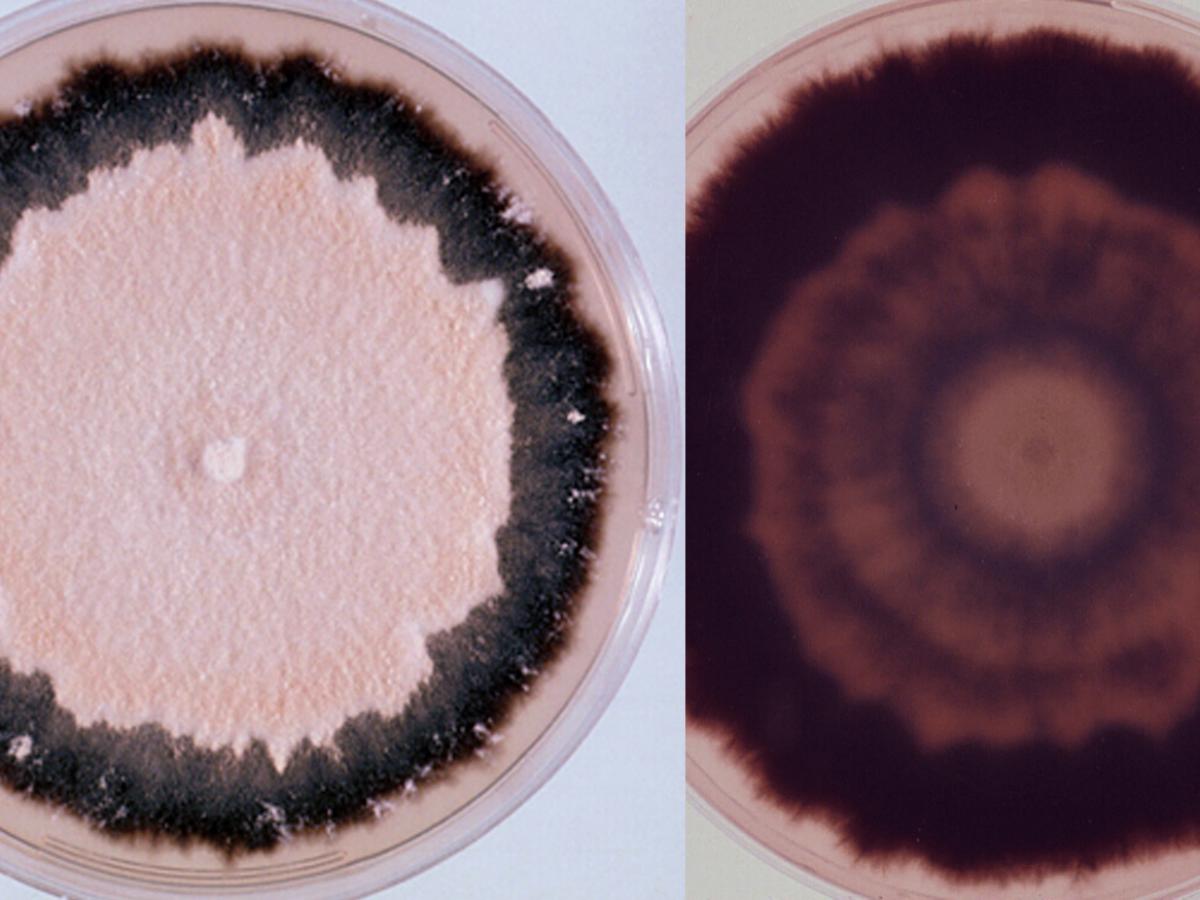
Arthroderma uncinatum culture.
Morphological description:
Colonies are usually flat, powdery, cream, tan to orange-tan in colour, with a blackish-purple submerged fringe and reverse. Macroconidia are numerous, smooth, thick-walled, elongate, cigar-shaped, 29-65 x 5-10 µm, and multiseptate with up to nine or ten septa. Microconidia are usually absent, but when present are ovate to pyriform in shape.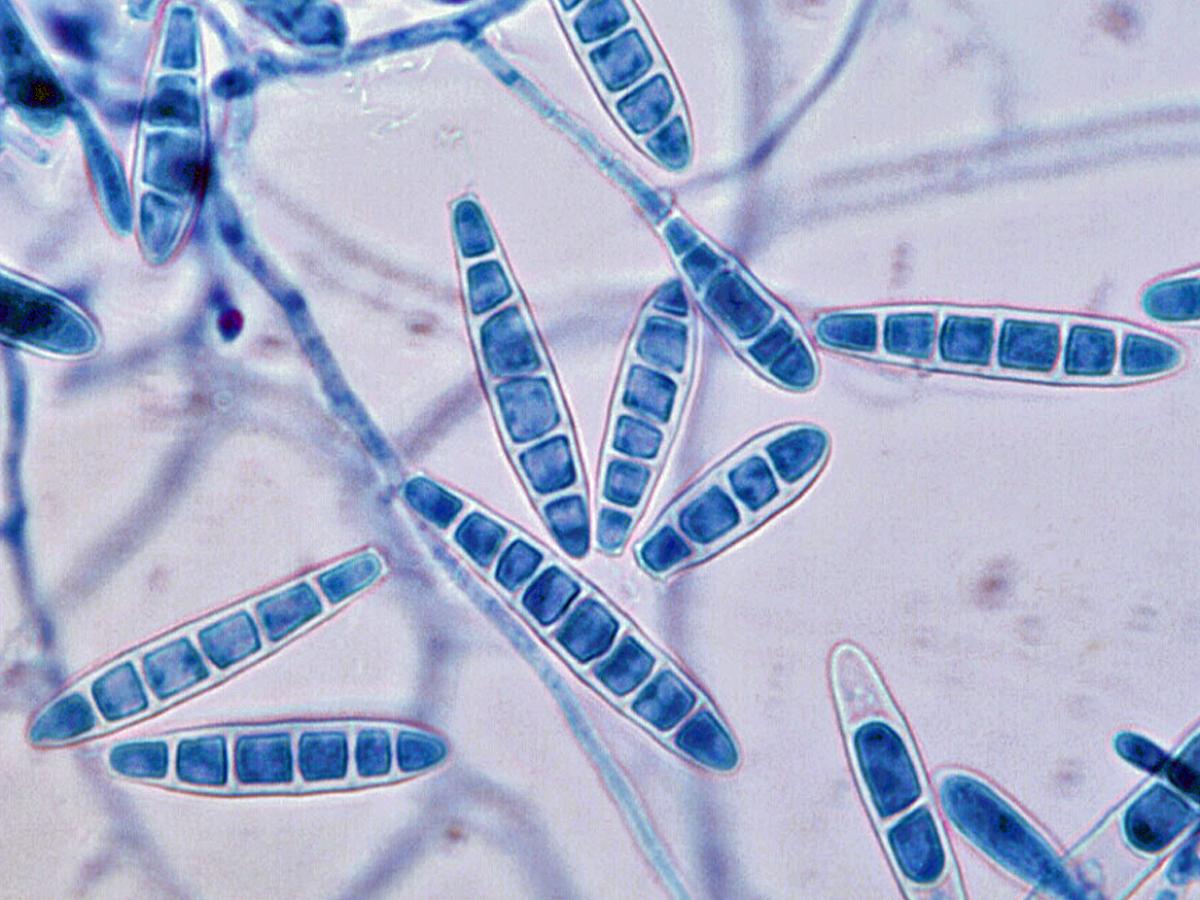
Arthroderma uncinatum macroconidia.
Key features:
Culture characteristics, macroconidial morphology, urease positive and good growth on Sabouraud’s 5% salt agar.Molecular identification:
ITS sequencing recommended (Gräser et al. 2008).References:
Rebell and Taplin (1970), Rippon (1988), de Hoog et al. (2015, 2016).
