Malassezia
Malassezia species are basidiomycetous yeasts and form part of the normal skin flora of humans and animals.
The genus now includes 16 species of which 15 are lipid dependent. These include M. arunalokei (human), M. caprae (goat, horse), M. cuniculi (rabbit), M. dermatis (human), M. equina (horse, cow), M. furfur (human, cow, elephant, pig, monkey, ostrich, pelican), M. globosa (human, cheetah, cow), M. japonica (human), M. nana (cat, cow, dog), M. obtusa (human), M. pachydermatis (dog, cat, carnivores, birds), M. restricta (human), M. slooffiae (human, pig, goat, sheep), M. sympodialis (human, horse, pig sheep), M. vespertilionis (bats) and M. yamatoensis (human) (Cabanes et al. 2011).
M. sympodialis, M. globosa, M. slooffiae and M. restricta are the most frequently found species responsible for colonisation of humans (Arendrup et al. 2014).
Malassezia species may cause various skin manifestations including pityriasis versicolor, seborrhoeic dermatitis, dandruff, atopic eczema and folliculitis. M. pachydermatis is known to cause external otitis in dogs. Fungaemia due to lipid-dependent Malassezia species usually occurs in patients with central line catheters receiving lipid replacement therapy, especially in infants (Tragiannides et al. 2010, Gaitanis et al. 2012, Arendrup et al. 2014).
Note: With the exception of M. pachydermatis, the primary isolation and culture of Malassezia species is challenging because in vitro growth must be stimulated by natural oils or other fatty substances. The most common method used is to overlay Sabouraud’s dextrose agar (SDA) containing cycloheximide (actidione) with olive oil or alternatively to use a more specialised media like modified Leeming and Notham agar (Kaneko et al. 2007), or modified Dixon’s agar (see culture techniques and media). However, CHROMagar Malassezia medium is now commercially available for the primary isolation and differentiation of the most common Malassezia species.
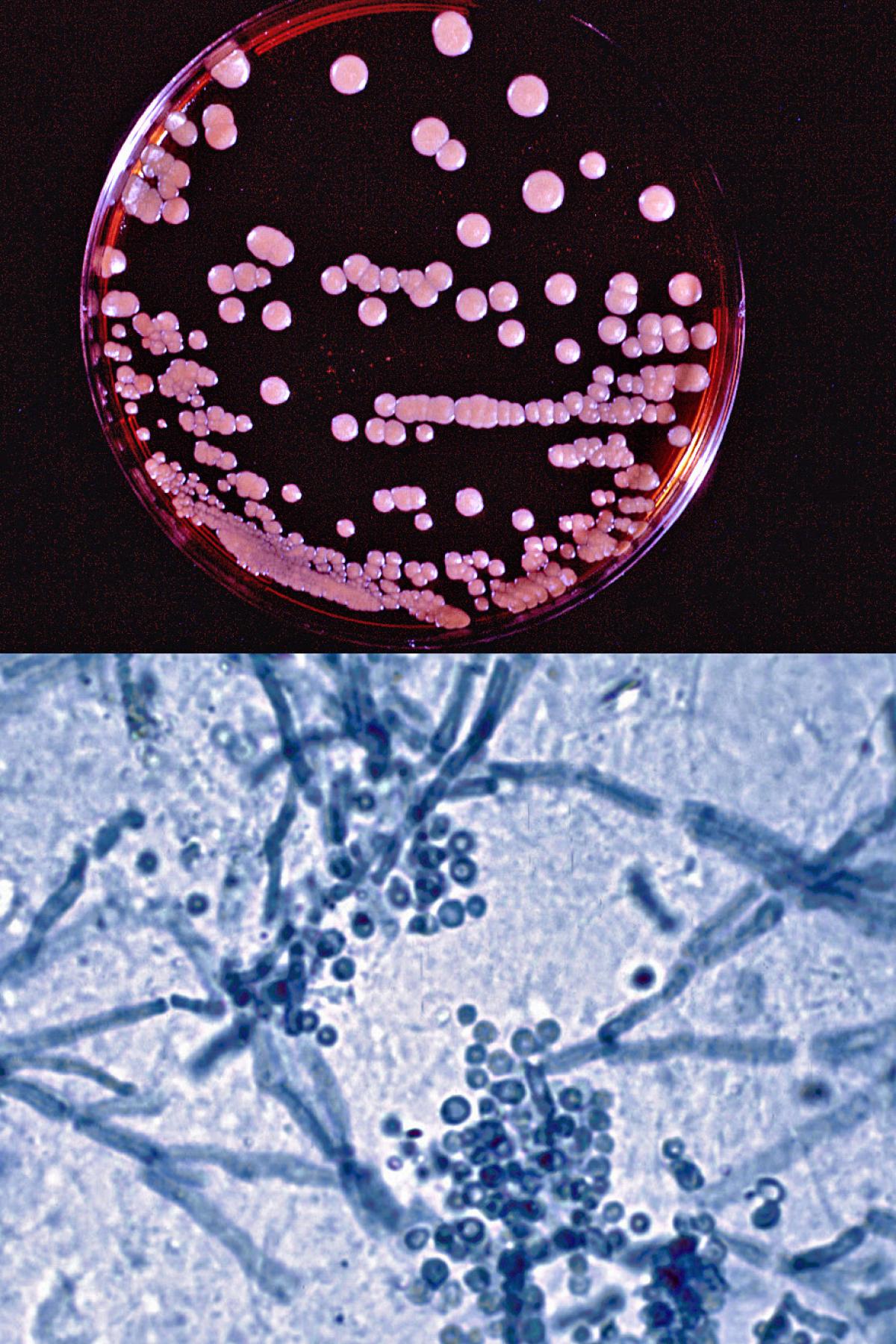
Malassezia furfur culture on modified Dixon's agar and direct microscopy of skin scrapings showing characteristic clusters of thick-walled round, budding yeast-like cells and short angular hyphal forms (the so called spaghetti and meatballs appearance) typically seen in pityriasis versicolor.
Comment:
For clinical management at the level of the individual patient, species identification is less important, although it is obviously needed for epidemiological surveillance and outbreak investigation (Arendrup et al. 2014).
RG-1 organisms.
Morphological description:
On media like modified Dixon’s agar, colonies are cream to yellowish, smooth or lightly wrinkled, glistening or dull, and with the margin being either entire or lobate. Malassezia is characterised by globose, oblong-ellipsoidal to cylindrical yeast cells. Reproduction is by budding on a broad base and from the same site at one pole (unipolar).
Molecular identification:
ITS and D1/D2 sequencing may be used for accurate species identification (de Hoog et al. 2015).
MALDI-TOF MS:
Capable of identifying all Malassezia species in concordance with those of ITS sequence analyses (Kolecka et al. 2014).
| Identification criteria for the differentiation of Malassezia species ((Cabanes et al., 2011, 2016; Honnavar et al., 2016; Lorch et al., 2018). + Positive, - Negative, v Variable, w Weak, s Slow, nd No Data, T = Tween | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Species | Buds | SDA | 40C | Cremophor EL |
T80 | T60 | T40 | T20 | Esculine | Catalase |
| M. arunalokei | narrow | - | - | - | v | - | - | - | nd | - |
| M. caprae | narrow | - | - | - | + | + | + | + | + | + |
| M. couiculi | narrow | - | + | - | - | - | - | - | nd | + |
| M. dermatis | wide | - | + | nd | + | + | + | + | nd | + |
| M. equina | narrow | - | - | - | + | + | + | + | + | + |
| M. furfur | wide | - | + | + | + | + | + | w | + | |
| M. globosa | narrow | - | - | - | - | - | - | - | + | |
| M. japonica | wide | - | - | nd | - | + | w | - | nd | + |
| M. nana | narrow | - | v | nd | w | + | + | v | nd | + |
| M. obtusa | wide | - | - | - | - | - | - | + | + | |
| M. pachydermatis | wide | + | + | -,w | + | + | -,w | v | v | |
| M. restricta | narrow | - | - | - | - | - | - | - | - | |
| M. slooffiae | wide | - | + | - | +,w | + | + | - | + | |
| M. sympodialis | narrow | - | + | + | +,w | + | + | + | + | + |
| M. vespertilionis | narrow | - | w | - | w | + | + | w | nd | - |
| M. yamatoensis | wide | - | - | nd | + | + | + | + | nd | + |
Antifungal susceptibility:
There is no standardised method for testing Malassezia species, special growth conditions are needed, and published results maybe variable. Note: Susceptibility testing is not recommended for guiding treatment.
Malassezia spp., limited data available (Velegraki et al., 2004; Miranda et al., 2007; Rojas et al., 2014); MIC µg/mL.
|
Antifungal |
M. furfur |
M. sympodialis |
M. globosa |
|||
|
Range |
MIC90 |
Range |
MIC90 |
Range |
MIC90 |
|
|
AmB |
0.125-16 |
2 |
0.06-4 |
2 |
0.03-4 |
1 |
|
FLU |
0.125-64 |
16 |
0.125-16 |
8 |
0.125-32 |
4 |
|
KETO |
0.03-4 |
0.25 |
0.03-0.25 |
0.125 |
0.03-0.5 |
0.06 |
|
ITRA |
0.03-0.25 |
0.25 |
0.03-0.125 |
0.06 |
0.03-0.125 |
0.125 |
|
VORI |
0.03-16 |
1 |
0.03-0.125 |
0.125 |
0.3-0. 25 |
0.125 |
|
POS |
0.03-32 |
2 |
0.03-0.06 |
0.03 |
0.03-0.06 |
0.06 |
References:
Guillot and Gueho (1995), Gueho et al. (1996), Guillot et al. (1996, 2000), Boekhout et al. (2010), Cafarchia et al. (2011), de Hoog et al. (2015).
